Begin data analysis in FlowJo with the user- friendly Graph Window. Quickly gate, navigate, and add statistics to your populations using FlowJo’s intuitive menu bars and icons.
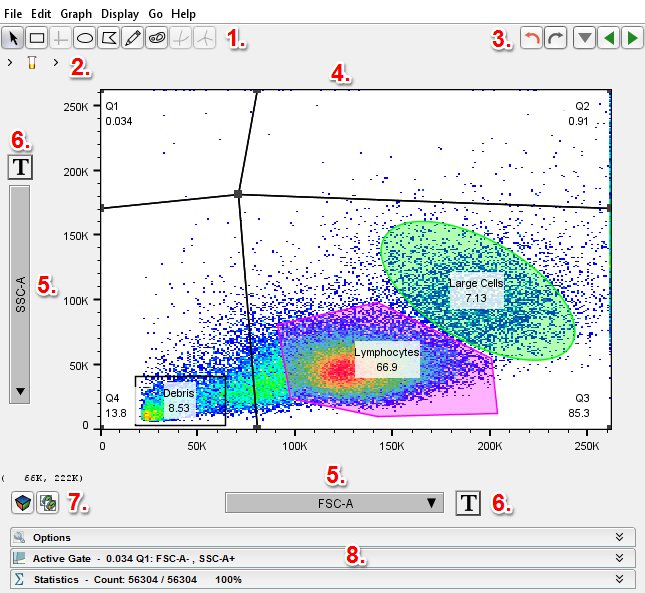
Data analysis in FlowJo begins with the Graph Window. The Graph Window provides an interface to begin gating your populations, adding statistics, and is the first step in creating your heirachical tree in the workspace. The Graph Window consists of several important components:
- The Gating Tools menu
- The Breadcrumb Bar
- The Undo/Redo and Navigation arrows
- The primary graphical display
- The X- and Y-axis drop down menus
- The Transform button
- The Duplicate Graph and 3D icons
- The Options, Active Gate, and Statistics expandable menus.
At the top of the window are the available tools for creating gates. For more information about the types of gates available, click here.
The Breadcrumb Bar provides a reference to where you are in your gating hierarchy, and has quick links to ancestral gates. For more information on the Breadcrumb Bar see below.
The primary graphical display has two drop-down menus (one for each axis) for viewing your cells under any acquired parameter (FITC, APC, PE, Cy5, FL-2, etc). A transform button sits adjacent to each axis drop-down menu to modify the axis scaling. For more information about axis transformation and scaling, click here.
At the bottom of the graph window are three expandable menus; Options, Active Gate, and Statistics. These menus allow you to change the type of graphical display (density plots, contour plots, histograms, etc.), the color, shading, and behavior of gates, and permit addition of a wide range of statistics.
Opening a Graph Window
To open a new Graph Window, double-click on any population node within the sample list in the workspace. Your cells will be displayed as events along an X and Y-axis (as part of a bivariate plot), allowing you to identify and gate different populations. By default, when you double-click on a node in your workspace, a graph window will open displaying the forward scatter-area (FSC-A) and the side scatter-area (SSC-A) on the X and Y-axis of the plot, respectively. This allows you to quickly gate out debris (dead cells, etc.) and target the overall cell populations you would like to analyze.
Gating Tools

Draw gates to create subsets of the data and select a variety of different representations of the data. For more info on gating and a list of gating tools click here.
The Breadcrumb Bar
The Breadcrumb Bar is a navigation tool that helps you toggle between and keep track of samples and subpopulations. The bar is located in the upper portion of the Graph Window and allows you to do three key things: It allows you to toggle between different samples within your workspace, helps you keep track of which subpopulations you are currently displaying and it allows you to jump rapidly to other subpopulations within the same gating tree.
For example, if you double click on a sample from your Workspace to open a Graph Window and create a lymphocyte gate (see image above), you will see a tube icon in the upper left-hand corner of the window. Clicking on the icon will open a drop-down menu that will show all of the other samples in the currently open Group in your Workspace. You can open a Graph Window with the same view and parameter set as the currently open Window by selecting one of these samples (below).
As you create further gates, such as a singlets gate and a CD4 subset gate, a list of your previous gates/subpopulations will be displayed next to the tube icon in the upper left-hand corner of the Graph Window. This list corresponds to the gating tree that is displayed in your Workspace under each sample.
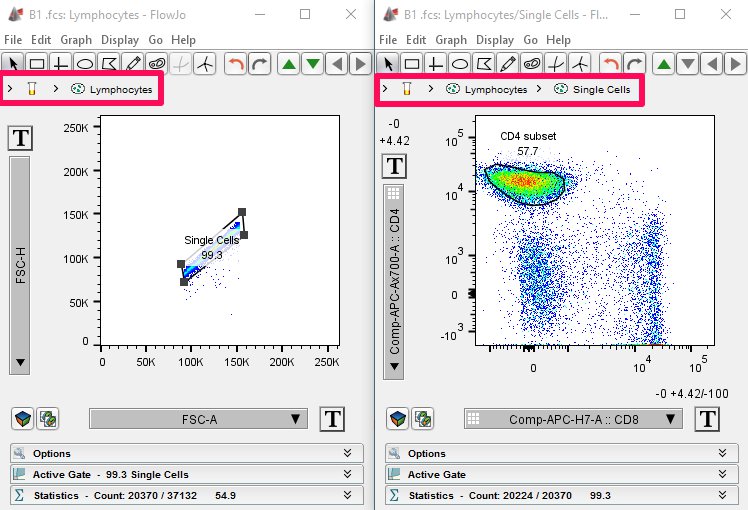
Finally, clicking on any population listed in the Breadcrumb Bar quickly opens a Graph Window for that population. This Graph Window will display the view and parameter set that was last displayed for the selected population.

Undo/Redo and Navigation arrows
To undo or redo an action (create/delete/modify a gate) click the Undo/Redo arrow button.

The navigation arrows allow you to Navigate the gating tree and sample set using arrow buttons.

These buttons allow for quick navigation among the available populations. The left and right arrows change the sample to previous or next, respectively, sample with an equal population/gate definition. The up arrow will open a graph window with the defining population (parent population).
If multiple graphs from the same sample are open, shift-click to change all open plots to next sample.
X & Y Axis Drop-down Menus

Use this menu (above) to select among the parameters available for display on the X & Y axis, you can also hold down the Alt key when clicking on the drop down to show uncompensated parameters. If a compensation matrix was applied to the population in the graph, those compensated parameters will also be available for selection. To view uncompensated parameters, click on the Display menu and select to Show Uncompensated Parameters. The Event Number parameter can be made available by selecting Show Event Number in the Display Menu.
Transform Button

Toggles between a linear, logarithmic, and biexponential scales. Use the transform button to change the scale type and add or subtract decades to help visualize and smooth your data.
3D Viewer and Duplicate Graph Window Icons

This icon opens the 3D viewer window, displaying the population on a 3 axis/parameter view in a 3D cube.
The duplicate graph window button opens a new graph window with the same view/parameter set.
Sometimes it may be helpful to view the same population in the Workspace hierarchy but with different parameters. By duplicating the graph window you can select different parameters and add additional gates and statistics to the original population of interest. For example, you may want to view the original lymphocytes subset in the CD3, CD4, CD45 and CD8 parameters. You could select CD3 (X) and CD4 (Y), then duplicate the graph and change the X and Y axes to CD45 and CD8 (or whichever combination you desire). This will allow you to view your population of interest with 3 or more parameters simultaneously.
Options, Active Gate, and Statistics menus

The first drop down menu (above), when clicked, expands additional graph options. Here, attributes of the current graph can be modified. The Type drop down menu allows to select form seven graph types contour plots, density plots, zebra plots, pseudocolor plots, dot plots, histograms, CDF plots. The Color boxes are used to change the colors of the graph. The left box changes the forground color, while the right box changes the background color. Select the Smooth checkbox to display certain graphs with a smooth color gradient for improved readability. By selecting the Show Outliers check box rare events are displayed as dots in a contour type graph. When the graph type is histogram the Y-Axis drop down menu allows for auto scaling or manual scaling of the Y-Axis. By default, this section is not expanded.
Active Gate Options
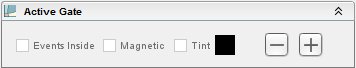
The second drop down menu (above), when clicked, expands options for the active gate in the graph window. When a gate is highlighted, relevant data appears to the right of this space. The Events Inside check box calculates the percent of events located inside the gate. Magnetic check box moves the gate until it accommodates that maximum number of events. The Tint check box allows for tinting the active gate by desired color. By default, this section is not expanded.
Generate Statistics
The third drop down menu (above), when clicked, expands statistics on the current population. Data related to the number of events gated, the total population, and frequency of total appear to the right of this space, as well as below it when revealed.
The Add Statistics button opens the Add Statistics window and allows for the selection of additional statistics. By default this section is not expanded.
Click on the link for more on Biexponential Transformation.
Click on the following link for detailed information on Exporting graph images for use in other programs.
Questions? Feel free to reach out to us at flowjo@bd.com
